Cell Division¶
On a 2D mesh¶
[1]:
import pandas as pd
import numpy as np
import json
import matplotlib.pylab as plt
import ipyvolume as ipv
%matplotlib inline
from tyssue import Sheet, config
from tyssue.geometry.planar_geometry import PlanarGeometry as geom
from tyssue.solvers.quasistatic import QSSolver
from tyssue.dynamics.planar_vertex_model import PlanarModel as model
from tyssue.draw import sheet_view
from tyssue.stores import load_datasets
from tyssue.topology.sheet_topology import remove_face, cell_division
[2]:
solver = QSSolver()
sheet = Sheet.planar_sheet_2d('division', 6, 6, 1, 1)
sheet.sanitize(trim_borders=True, order_edges=True)
geom.update_all(sheet)
sheet.get_opposite()
# ## Set up the model
nondim_specs = config.dynamics.quasistatic_plane_spec()
dim_model_specs = model.dimensionalize(nondim_specs)
sheet.update_specs(dim_model_specs, reset=True)
print("Number of cells: {}\n"
" edges: {}\n"
" vertices: {}\n".format(sheet.Nf, sheet.Ne, sheet.Nv))
# ## Minimize energy
res = solver.find_energy_min(sheet, geom, model)
# ## View the result
draw_specs = config.draw.sheet_spec()
draw_specs['vert']['visible'] = False
draw_specs['edge']['head_width'] = 0.1
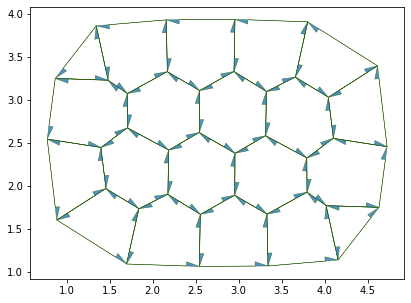
fig, ax = sheet_view(sheet, **draw_specs)
fig.set_size_inches(12, 5)
Reseting column is_alive of the face dataset with new specs
Reseting column ux of the edge dataset with new specs
Reseting column uy of the edge dataset with new specs
Reseting column is_active of the vert dataset with new specs
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
Number of cells: 20
edges: 100
vertices: 38
[3]:
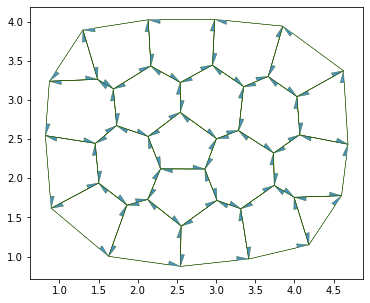
daughter = cell_division(sheet, 7, geom, angle=np.pi/2)
res = solver.find_energy_min(sheet, geom, model)
print(res['success'])
fig, ax = sheet_view(sheet, **draw_specs)
fig.set_size_inches(12, 5)
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
True
Division in a 3D single layer epithelium¶
[4]:
from tyssue.io.hdf5 import save_datasets, load_datasets
# redefine cell_division from monolayer related topology module
from tyssue.topology.monolayer_topology import cell_division
from tyssue.core.monolayer import Monolayer
from tyssue.geometry.bulk_geometry import ClosedMonolayerGeometry as monolayer_geom
from tyssue.dynamics.bulk_model import ClosedMonolayerModel
from tyssue.draw import highlight_cells
[5]:
datasets = load_datasets('data/small_ellipsoid.hf5',
data_names=['vert', 'edge',
'face', 'cell'])
specs = config.geometry.bulk_spec()
monolayer = Monolayer('ell', datasets, specs)
monolayer_geom.update_all(monolayer)
specs = {
"edge": {
"line_tension": 0.0,
},
"face": {
"contractility": 0.01,
},
"cell": {
"prefered_vol": monolayer.cell_df['vol'].median(),
"vol_elasticity": 0.1,
"prefered_area": monolayer.cell_df['area'].median(),
"area_elasticity": 0.1,
},
"settings": {
'lumen_prefered_vol': monolayer.settings['lumen_vol'],
'lumen_vol_elasticity': 1e-2
}
}
monolayer.update_specs(specs, reset=True)
res = solver.find_energy_min(monolayer, monolayer_geom, ClosedMonolayerModel)
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
Reseting column line_tension of the edge dataset with new specs
Reseting column contractility of the face dataset with new specs
Reseting column prefered_vol of the cell dataset with new specs
Reseting column vol_elasticity of the cell dataset with new specs
[6]:
mother = 8
daughter = cell_division(monolayer, mother,
orientation='vertical')
monolayer.validate()
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
[6]:
True
[7]:
rho = np.linalg.norm(monolayer.vert_df[monolayer.coords], axis=1)
draw_specs['edge']['color'] = rho
draw_specs['face']['visible'] = True
ipv.clear()
highlight_cells(monolayer, mother, reset_visible=True)
fig, mesh = sheet_view(monolayer, mode="3D",
coords=['z', 'x', 'y'], **draw_specs)
fig
[8]:
mother = 18
daughter = cell_division(monolayer, mother,
orientation='horizontal')
monolayer.validate()
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
[8]:
True
[9]:
rho = np.linalg.norm(monolayer.vert_df[monolayer.coords], axis=1)
draw_specs['edge']['color'] = rho
draw_specs['face']['visible'] = True
ipv.clear()
highlight_cells(monolayer, mother)
fig, mesh = sheet_view(monolayer, mode="3D",
coords=['z', 'x', 'y'], **draw_specs)
fig
Energy minimisation of the monolayer after division¶
[10]:
monolayer.cell_df.loc[[mother, daughter], 'prefered_area'] /= 2
monolayer.cell_df.loc[[mother, daughter], 'prefered_vol'] /= 3
monolayer.settings['lumen_prefered_vol'] = monolayer.settings['lumen_vol']
monolayer.settings['lumen_vol_elasticity'] = 1e-2
res = solver.find_energy_min(monolayer, monolayer_geom, ClosedMonolayerModel)
print(res['message'])
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
b'CONVERGENCE: NORM_OF_PROJECTED_GRADIENT_<=_PGTOL'
[11]:
rho = np.linalg.norm(monolayer.vert_df[monolayer.coords], axis=1)
draw_specs['edge']['color'] = rho
ipv.clear()
highlight_cells(monolayer, mother)
fig, mesh = sheet_view(monolayer, mode="3D",
coords=['z', 'x', 'y'], **draw_specs)
fig
[12]:
from tyssue.generation import three_faces_sheet, extrude
from tyssue.geometry.bulk_geometry import MonoLayerGeometry
datasets_2d, _ = three_faces_sheet(zaxis=True)
datasets = extrude(datasets_2d, method='translation')
eptm = Monolayer('test_volume', datasets,
config.geometry.bulk_spec(),
coords=['x','y','z'])
#eptm.vert_df[eptm.coords] += np.random.normal(scale=1e-3,
# size=eptm.vert_df[eptm.coords].shape)
MonoLayerGeometry.update_all(eptm)
for orientation in ['vertical', 'horizontal']:
print(orientation)
daughter = cell_division(eptm, 1,
orientation=orientation)
eptm.reset_topo()
eptm.reset_index()
MonoLayerGeometry.update_all(eptm)
print(f'Valid division for {orientation}:')
print(eptm.validate()) #break
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
vertical
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
Valid division for vertical:
True
horizontal
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
/home/docs/checkouts/readthedocs.org/user_builds/tyssue/conda/stable/lib/python3.7/site-packages/tyssue/utils/utils.py:35: FutureWarning: Support for multi-dimensional indexing (e.g. `obj[:, None]`) is deprecated and will be removed in a future version. Convert to a numpy array before indexing instead.
df_nd = df[:, None]#np.asarray(df).repeat(ndim).reshape((df.size, ndim))
Valid division for horizontal:
True
[ ]: